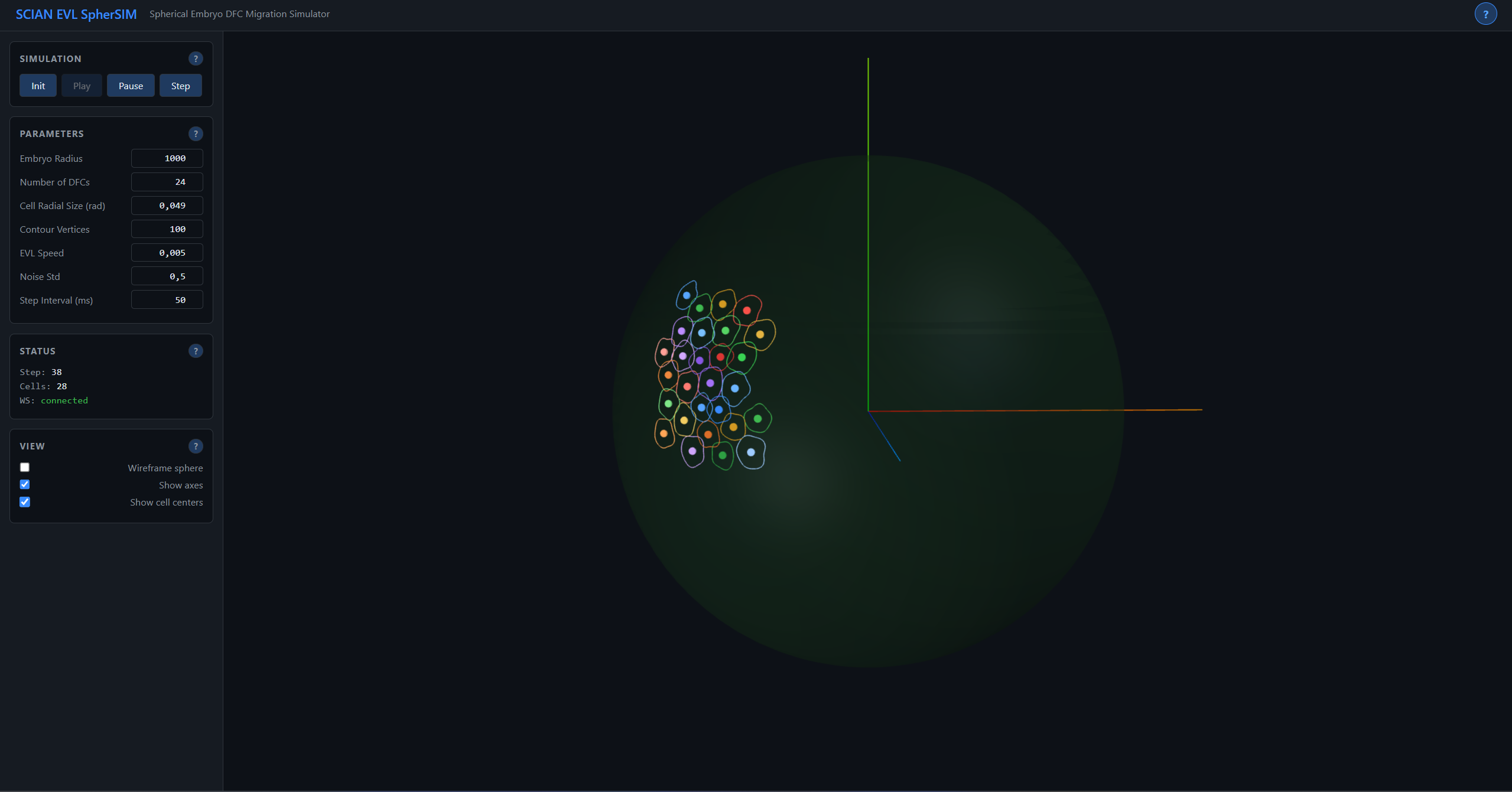
3D Embryo Cell Migration Simulator
Published:
A 3D simulation of Deep Forming Cell (DFC) collective migration on the surface of a spherical zebrafish embryo during epiboly — a fundamental process in vertebrate development where cells spread to cover the yolk cell.
Biological context
During zebrafish gastrulation, the Enveloping Layer (EVL) expands from the animal pole toward the vegetal pole. Deep Forming Cells (DFCs) are carried along by the advancing EVL margin, eventually clustering to form the Kupffer’s vesicle — an organ critical for establishing left-right body asymmetry. Understanding the mechanics of this collective migration requires modeling how cells move, collide, and respond to tissue-level forces on a curved surface.
The simulation
Cells move on the surface of a sphere using an AER (Azimuth-Elevation-Radius) coordinate system with automatic wrapping at the poles:
- Azimuth: horizontal angle (-pi to pi), analogous to longitude, with periodic wrapping
- Elevation: vertical angle (-pi/2 to pi/2), analogous to latitude, clamped at the poles
- Radius: constant (cells remain on the sphere surface)
Each cell’s contour is discretized as N vertices: vertex[i] = (center_az + rcos(2pii/N), center_el + rsin(2pii/N), R_embryo). Cell velocity has a deterministic component from EVL coupling (base_velocity = [0, -(pi/2)*evl_speed, 0]) plus a Gaussian stochastic term scaled by migration speed, capturing the biological variability of cytoskeletal dynamics. Pairwise collision detection in angular space (angular_dist < size_a + size_b) with symmetric push-apart resolution ensures cells maintain physical separation while migrating collectively.
Technical highlights
- Backend: Python/FastAPI with real-time WebSocket streaming of simulation state
- Frontend: Three.js (r128) for interactive 3D visualization with orbit controls
- Cartesian push collision model: improved cell-cell interactions using 3D displacement vectors projected back onto the sphere surface
- Layer-based DFC management: cells organized by developmental layer with configurable coupling forces
- Controls: Play/Pause/Step for continuous or frame-by-frame exploration
- Configurable: Cell count, radial size, EVL speed, noise amplitude, collision parameters
- Legacy: Preserves the original MATLAB implementation alongside the modern version
- 23+ automated tests across cell, collision, geometry, and simulation modules
From SCIAN-Lab
This project originated at the Laboratory for Scientific Image Analysis (SCIAN-Lab) at the Universidad de Chile, where I worked on computational tools for developmental biology research in collaboration with the NEMO Millennium Nucleus.
Live application
▶ Live demo — sphersim.fasl-work.com — 3D embryo cell migration simulator on a spherical surface.