Cellular Potts Model Simulator
Published:
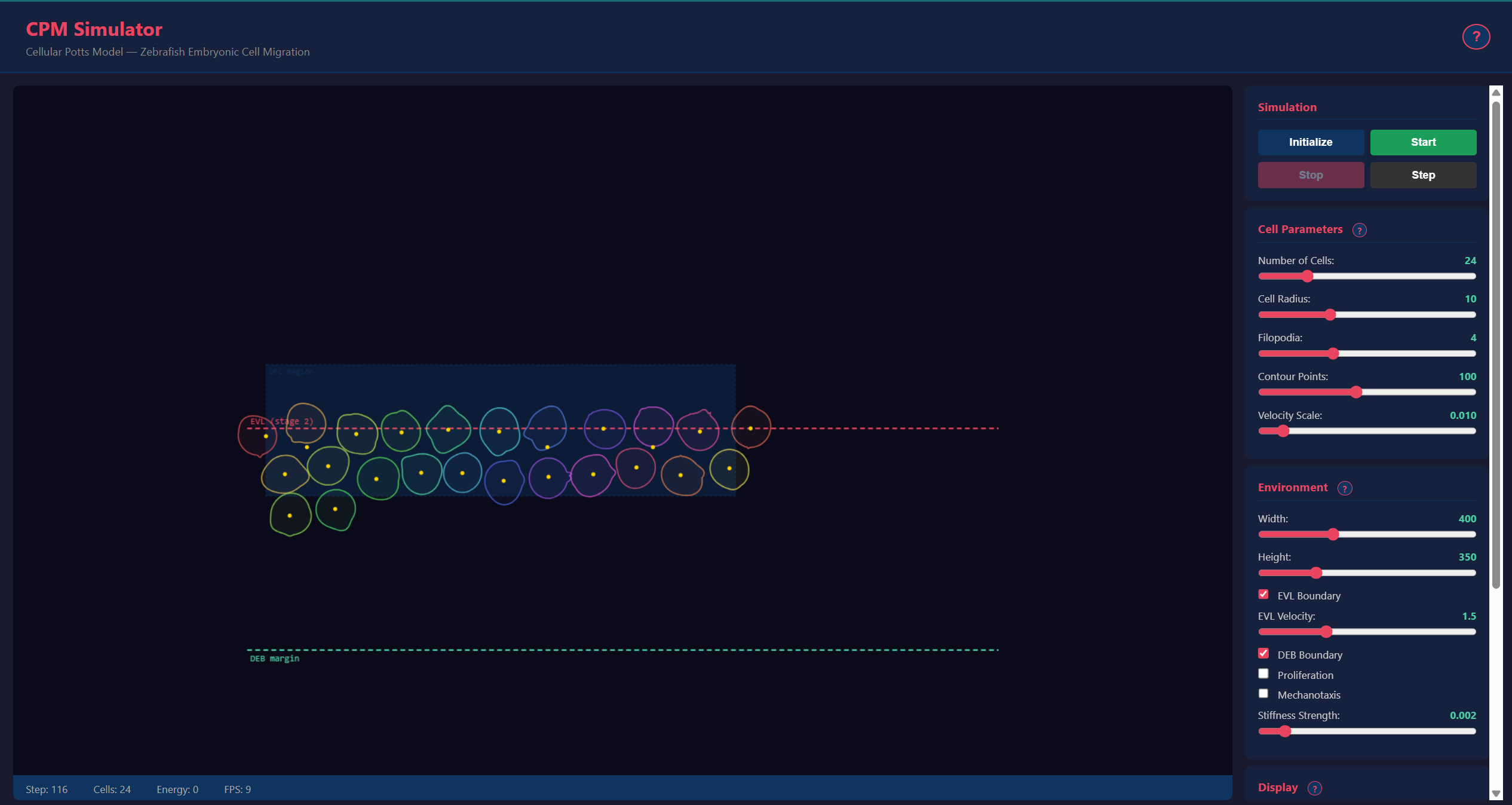
A Cellular Potts Model (CPM) simulator for modeling collective migration of Dorsal Forerunner Cells (DFCs) during zebrafish Kupffer’s vesicle formation. Unlike particle-based models, the CPM represents cells as deformable bodies on a lattice, capturing shape changes, area constraints, and contact interactions naturally.
Why a Cellular Potts Model?
Real cells are not point particles. They have area, they deform when they push against neighbors, and they extend structures (filopodia) that generate directional forces. The CPM captures all of this on a lattice where each cell occupies multiple pixels, and the system evolves by minimizing a Hamiltonian that balances:
- Area constraint: cells resist compression and expansion
- Perimeter constraint: cells resist excessive deformation
- Adhesion energy: cells stick to neighbors with type-dependent strength
- Motility: Gaussian filopodia generate directed forces toward migration targets
Gaussian filopodia model
Each cell’s radial contour is defined by a set of filopodial protrusions modeled as Gaussian bumps on a base radius:
R(theta) = max over j of { R0 + A_j * exp( -(theta - theta0_j)^2 / (2 * W_j^2) ) }
where R0 is the resting cell radius, A_j is the amplitude (extension length), theta0_j is the angular direction, and W_j is the angular spread of filopodium j. The cell’s velocity is then derived from the shape asymmetry of its membrane contour – a cell with an asymmetric protrusion migrates in that direction. Filopodium angles undergo a Brownian random walk at each time step, producing realistic exploratory motility. A two-pass collision resolution (soft forward pass + hard backward pass) ensures stable cell-cell separation without oscillation.
Key features
- Deformable cell model with configurable Gaussian filopodia
- Durotaxis mechanics: cells respond to stiffness gradients in the tissue, biasing migration toward stiffer regions — a key mechanism in developmental biology
- Real-time 2D Canvas visualization at 10-50 FPS
- Per-cell color coding for tracking individual cells
- Tissue boundary dynamics: EVL epiboly front and DEB convergence margin
- Cell proliferation with configurable 8-stage schedule
- Two-pass collision resolution (soft + hard pass)
- REST API with Swagger/ReDoc documentation
- ~30 automated tests covering cells, agents, and simulation logic
Technical stack
- Backend: Python/FastAPI with NumPy for lattice operations
- Frontend: HTML5 Canvas 2D rendering with interactive slider controls
- Communication: REST API + WebSocket for live simulation streaming
- Legacy: Previous C++/CLI codebase (2014-2024) preserved for reference
Research context
Developed during my work at SCIAN-Lab and the BNI at the Universidad de Chile, this simulator supported research in developmental biology, helping understand the physical mechanisms behind collective cell migration in vertebrate embryos.
Live application
▶ Live demo — cpm.fasl-work.com — Cellular Potts Model simulator — zebrafish DFC migration.